1BIOPAT.Biopatologia Molecular, Barcelona, Spain; 2Hospital de Barcelona, Barcelona, Spain; 3Histopat Laboratoris, Barcelona, Spain; 4Statistics Department, Universitat de Barcelona, Barcelona, Spain; 5Hospital Universitari de Bellvitge, Barcelona, Spain; 6Hospital del Mar, Barcelona, Spain; 7Hospital Mútua de Terrassa, Terrasa, Spain; 8Hospital General de Vic, Vic, Spain and 9Hospital Plató, Barcelona, Spain.
Abstract
High microsatellite instability (MSI-H) allows the identification of a subset of colorectal carcinomas associated with good prognosis and a higher incidence of Lynch syndrome. The aim of this work was to assess the interobserver variability and optimize our MSI-H prediction model previously published based on phenotypic features. The validation series collected from five different hospitals included 265 primary colorectal carcinomas from the same number of patients. The eight clinicopathological parameters that integrate our original model were evaluated in the corresponding centers. Homogeneity assessment revealed significant differences between hospitals in the estimation of the growth pattern, presence of Crohn-like reaction, percentage of cribriform structures, and Ki-67 positivity. Despite this observation, our model was globally able to predict MSI-H with a negative predictive value of 97.0%. The optimization studies were carried out with 615 cases and resulted in a new prediction model RERtest8, which includes the presence of tumor infiltrating lymphocytes at the expense of the percentage of cribriform structures. This refined model achieves a negative predictive value of 97.9% that is maintained even when the immunohistochemical parameters are left out, RERtest6. The high negative predictive value achieved by our models allows the reduction of the cases to be tested for MSI to less than 10%. Furthermore, the easy evaluation of the parameters included in the model renders it a useful tool for the routine practice and can reinforce other published models and the current clinical protocols to detect the subset of colorectal cancer patients bearing hereditary nonpolyposis colorectal cancers risk and/or MSI-H phenotype.
Keywords Microsatellite instability – Prediction model – Colorectal cancer – Pathological parameters – Hereditary nonpolyposis colorectal cancer
Introduction
The study of global genomic and epigenomic changes occurring during colorectal carcinogenesis, mainly chromosomal instability (CIN), microsatellite instability (MSI), and CpG island methylation phenotype, is allowing the classification of colorectal tumors according to their molecular status
The most common CIN pathway, involving approximately 90% of sporadic colorectal tumors and all cases of familial adenomatous polyposis, follows the adenoma carcinoma sequence through sequential accumulation of a series of mutations at crucial regulatory genes showing frequent chromosomal gains and losses [3]. On the contrary, tumors developing through the MIN pathway show a high degree of MSI (MSI-H) [4] which is due to alterations in the mismatch repair genes (MMR), mainly MLH1, MSH2, and MSH6 [5]. These alterations can be caused by germline mutations, which give rise to hereditary nonpolyposis colorectal cancers (HNPCC) [6] or by epigenetic silencing through hypermethylation of the MLH1 promoter [7]. Patients carrying both sporadic and hereditary MIN colorectal cancers exhibit a better prognosis and a poor response to 5-fluorouracil-based chemotherapy compared to those with CIN tumors [8, 9]. It is therefore essential to distinguish between these two main types of carcinomas.
Testing for MSI or for loss of MMR in all colorectal cancer cases would be a very expensive and time-consuming task to identify only the approximately 10% of tumors following the MIN pathway. It has become clear during the last years that these tumors display common morphological characteristics such as proximal location, mucinous differentiation, solid growth pattern, presence of intraepithelial lymphocytes and Crohn’s-like lymphocytic reaction which can make them distinguishable from CIN tumors [10-12].
Several studies have been reported in the literature in an effort to identify MIN status of colorectal tumors. Recently, Jenkins and colleagues [13] used pathology features described in the Bethesda guidelines to predict MSI in a group of patients diagnosed with colorectal carcinomas before age 60 with the primary aim of identifying HNPCC candidates. Also lately another model has been published by Greenson and colleagues [14] which classifies MSI-H tumors with an 85% accuracy looking at pathologic features. The objective of our work was to validate and improve with a multicentric study a logistic model based on clinicopathological features previously designed in our laboratories to predict microsatellite instability [15]. We used a nonselected population of 615 colorectal carcinomas to provide pathologists with an easy to use tool to identify a large subset of carcinomas which do not present a MSI-H status and would need no MIN analysis. The major aim of our study was to achieve a high negative predictive value which would reduce greatly the number of cases to be tested for MSI.
Materials and methods
Validation series
For our validation series, a total of 265 unselected primary colorectal cancer cases were prospectively collected from five different centers in the area of Catalonia, Spain.
Cases were evaluated by the corresponding pathologist following standard published criteria without previous specific training. Immunohistochemical analysis was also carried out at each participating center according to their own routine protocols in order to preserve interobserver heterogeneity. Histopathologic features were recorded as previously described [15]. Briefly such features included categorical and numerical variables. Categorical variables recorded were gender, tumor location, tumor configuration, extent of invasion, intramural and extramural thin-walled vessel invasion (TWVI), venous vessel invasion (VVI), perineural invasion (PNI), growth pattern, peritumoral Crohn-like reactivity which was considered positive when at least three nodular aggregates of lymphocytes were present within a single low power field (4× magnification), presence of tumor infiltrating lymphocytes (TIL) characterized by the finding of at least four intraepithelial lymphocytes in a high power field (40× magnification) [16], and presence of residual adenomas. Numerical variables evaluated were age, tumor size, number of affected lymph nodes, percentage of solid [17], mucinous, cribriform, micropapillary and microglandular patterns, as well as expression of Ki-67 and p53 by immunohistochemistry. Representative formalin-fixed paraffin-embedded blocks of paired tumor and normal tissue were then sent to our center for MSI-H prediction and MSI-H molecular analysis.
Simultaneously a prospective series of 148 cases was collected and identically assessed in our institution; this added interobserver variability and also serve to increase the global validation series.
Optimization series
The global series employed for the improvement of our prediction model included the 265 external cases from our validation series, another 146 from the prospective series evaluated in our institution, plus the 204 cases from our first study [15]. A summary of the histopathological variables of our global series is displayed on Table 1.
Table 1 Clinicopathological variables of the global optimization series
| Categorical variable | MSS (%) n = 563 | MSI-H(%) n=52 |
| Gender | ||
| Male | 336 (59.7) | 22 (42.3) |
| Female | 227 (40.3) | 30 (57.7) |
| Location | ||
| Proximal | 177 (31.4) | 45 (86.5) |
| Distal | 386 (68.6) | 7 (13.5) |
| Configuration | ||
| Exophytic | 244 (43.3) | 23 (44.2) |
| Ulcerated | 230 (40.9) | 23 (44.2) |
| Stenosing | 89 (15.8) | 6 (11.5) |
| Extent of invasion (pT) | ||
| pT1 | 37 (6.6) | 1 (1.9) |
| pT2 | 83 (14.7) | 7 (13.5) |
| pT3 | 271 (48.1) | 33 (63.5) |
| pT4 | 172 (30.6) | 11 (21.2) |
| Intramural TWVI | ||
| Present | 172 (30.6) | 16 (30.7) |
| Absent | 391 (69.4) | 36 (69.2) |
| Extramural TWVI | ||
| Present | 141 (25.0) | 11 (21.2) |
| Absent | 422 (75.0) | 41 (78.9) |
| Intramural VVI | ||
| Present | 40 (7.1) | 2 (3.8) |
| Absent | 522 (92.7) | 50 (96.2) |
| Extramural VVI | ||
| Present | 113 (20.1) | 8 (15.4) |
| Absent | 450 (79.9) | 44 (84.6) |
| Intramural PNI | ||
| Present | 54 (9.6) | 2 (3.8) |
| Absent | 509 (90.4) | 50 (96.2) |
| Extramural PNI | ||
| Present | 76 (13.5) | 3 (5.8) |
| Absent | 487 (86.5) | 49 (94.2) |
| Growth pattern | ||
| Expansive | 193 (34.3) | 32 (61.5) |
| Infiltrative | 370 (65.7) | 20 (38.5) |
| Crohn-like lymphoid reaction | ||
| Present | 190 (33.7) | 41 (78.8) |
| Absent | 373 (66.3) | 11 (21.2) |
| TIL | ||
| Present | 86 (15.3) | 29 (55.8) |
| Absent | 477 (84.7) | 23 (44.2) |
| Residual adenoma | ||
| Present | 176 (31.3) | 15 (28.8) |
| Absent | 387 (68.7) | 37 (71.2) |
| Numerical variable | Mean ± SD | Mean ± SD |
| Age | 69.1 ± 11.6 | 69.8 ± 12.9 |
| Tumor size (mm) | 43.2 ± 21.1 | 57 .5± 17.1 |
| Solid pattern (%) | 5.0 ± 12.9 | 23.7 ± 35.2 |
| Mucinous pattern (%) | 9.2 ± 20.7 | 32.3 ± 32.5 |
| Cribiform pattern (%) | 4.3 ± 10.1 | 12.3 ± 19.7 |
| Micropapillary pattern (%) | 2.0 ± 6.9 | 0.3 ± 1.5 |
| Microglandular pattern (%) | 3.2 ± 9.8 | 0.6 ± 2.2 |
| Nodal involvement (n) | 2.0 ± 3.7 | 2.2 ± 5.7 |
| Ki67 proliferative index (%) | 62.4 ± 21.7 | 72.6 ± 17.0 |
| p53 overexpression (%) | 43.3 ± 38.5 | 17.9 ± 24.1 |
Microsatellite instability analysis
Genomic DNA was extracted from ten 5-µm-thick sections of paired normal and tumor samples by macrodissection of selected areas followed by a proteinase K-phenol/chloroform protocol. DNA (200 ng) were used for each specific PCR reaction after the assessment of DNA quality.
MSI status was evaluated as described previously [15], using a panel of 11 microsatellites composed by the five microsatellites from the NCI panel (BAT25, BAT26, D5S346, D2S123, and D17S250) [18] in a multiplex PCR reaction, five additional microsatellites originally aimed at detecting the LOH status of chromosome 18q (D18S55, D18S58, D18S61, D18S64, and D18S69) [19] also amplified in a multiplex reaction and a microsatellite at the TP53 locus on 17p (P53CA) [20]. Fluorescent amplicons were analyzed on an automated ABI PRISM® 310 Genetic Analyzer using the GeneScan software (PE Applied Biosystems). According to the consensus definitions of the US NCI, tumors were classified as exhibiting MSI-H when 30% or more of the tested loci resulted unstable and non-MSI-H when they were less than 30%. Tumors exhibiting low microsatellite instability (up to three unstable markers out of 11) were considered together with stable tumors.
Immunohistochemical analysis
Immunohistochemical analysis in our institution was performed by ABC immunoperoxidase staining method, using mouse monoclonal antibodies DO-7 and MIB-1 (DakoCytomation, Denmark A/S) detecting p53 and Ki-67 proteins, respectively. Positive and negative controls were included in each experiment. Immunohistochemical evaluation was conducted double-blind by scoring the estimated percentage of tumor cells showing nuclear staining.
Statistical analysis
Our previous experience with the logistic regression model [15] capable to predict very accurately MSI instability impels us to assay again this modeling approach with the multicenter data set. Our purpose is to obtain a mathematical expression in order to estimate the probability of a tumor exhibiting MSI-H according to the following equation:
(*)
Where P is the expected probability of MSI-H and
is the development of the linear component of (*). β 0 is the independent term of the regression equation, β i is the regression coefficient for the i-th explanatory variable, and x i is the value of the i-th variable for any individual tumor. Notice that for dichotomous variables, x i assumes value 1 or 0.
Instead of the classical stepwise regression used in our previous publication [15] we follow here a different strategy to select the variables included in the model and to estimate their coefficients. In a first stage of our current approach, we use an automatic selection procedure of the variables based on the recent developments of the statistical topic known as shrinkage methods for model selection. More precisely, we use the methodology Regularization Paths for Generalized Linear Models via Coordinate Descent described in Friedman et al. [21, 22] implemented by these authors in the glmnet package running on the R statistical environment [23]. The glmnet package does not currently provide any stopping criteria to the user, we employ in our R script the Schwartz criteria combined with the tenfold cross-validation technique. For each of the ten validation sets, where it is successively excluded 10% of the full data set, the glmnet package with the Schwartz criteria [24] select the subset of variables that must be introduced in the logistic equation among our set of 30 maximum possible explanatory candidates. Therefore, a table with the frequency of each of the possible explanatory variable included in the ten final models, one for each validation data set, can be obtained.
These results are used in the second stage of our approach, which discards, if any, variables included in less than 70% of the final models. In order to configure the final form of the equation, we take into account in this step the clinicopathological knowledge of the variables.
When the variables in the right side of the equation are definitively established, we proceed to estimate the coefficients in the logistic model and validate its predictive capability. In this third stage, we use a huge resampling approach, where the basic element consists on a random split of the full data set in two subsets. The first subset, the training set, is used to compute the coefficients of the logistic model. They are obtained also via glmnet package, with the shrinkage parameter equal to 0. Then, the predictive capability is computed on the second subset, the validation set. We repeat 1,000 times this basic procedure of random split and predictive computations on the resulting validation set in order to avoid potential biases on the results due to the random selection. We compute the average value of the coefficients and the average prediction capability of the new proposed model over the random 1,000 data sets. We compare its results to the obtained with our previous prediction model [15] using the same 1,000 validation data sets.
Results
Validation studies
The first approach to test the robustness of our original prediction model was the assessment of the homogeneity between the participating hospitals, following the model’s equation:
Cases were assigned as MSI-H when P > 0.29. The coefficients of the linear component (x) corresponded to: (a) location (0 = proximal, 1 = distal), (b) growth pattern (0 = expansive, 1 = infiltrative), (c) Crohn-like response (0 = present, 1 = absent), (d) solid pattern%, (e) mucinous pattern%, (f) cribriform pattern, (g) Ki-67%, and (h) p53 accumulation%.
Significant differences were revealed in the estimation of the expansive growth, presence of Crohn-like inflammatory response, percentage of cribriform structures, and Ki-67 expression (Table 2). Despite this interobserver discrepancies, our original model was globally able to predict MSI-H with a negative predictive value of 97.0%, reducing the number of cases to be tested for MSI to just 11%. The results obtained with our own prospective series were equivalent, achieving a negative predictive value of 97.8% and did not differ from those obtained in the initial study where the negative predictive value was 97.8%. Accuracy, sensitivity, specificity, positive, and negative predicted values of the three series are shown in Table 3.
Table 2 Homogeneity assessment between the five participating hospitals
| Center 1 | Center 2 | Center 3 | Center 4 | Center 5 | p value | |
| Proximal | 6 | 12 | 18 | 42 | 26 | 0.33 |
| Distal | 4 | 15 | 42 | 58 | 42 | |
| Infiltrative | 6 | 4 | 37 | 44 | 26 | <0.001 |
| Expansive | 4 | 23 | 23 | 56 | 50 | |
| Crohn-like present | 1 | 11 | 15 | 69 | 17 | <0.001 |
| Crohn-like absent | 9 | 16 | 45 | 31 | 51 | |
| Solid pattern (%) | – | 5.7 | 1.2 | 1.2 | 6.5 | 0.20 |
| Mucinous pattern (%) | – | 11 | 10 | 10 | 6.5 | 0.81 |
| Cribiform pattern (%) | – | 25 | 17 | 11 | 18 | <0.001 |
| Ki-67 proliferative index (%) | – | 68 | 69 | 67 | 78 | 0.004 |
| p53 expression (%) | – | 40 | 55 | 43 | 47 | 0.32 |
Significant differences are highlighted in bold
Table 3 Statistical parameters achieved in our initial study, validation set, and prospective set
| Initial study | Validation set | Our Prospective set | |
| Accuracy | 95.10 | 91.32 | 91,41 |
| Sensitivity | 77.78 | 66.67 | 83.33 |
| Specificity | 96.77 | 93.44 | 92.41 |
| Positive predictive value | 70.00 | 46.67 | 57.69 |
| Negative predictive value | 97.83 | 97.02 | 97.81 |
| Total (n) | 204 | 265 | 148 |
Optimization studies
Out of the 615 colorectal carcinomas included in our global series, 52 (8.5%) exhibited MSI-H. The clinicopathological variables of this series are illustrated in Table 1. The glmnet cross-validation analysis carried out with the global series, involving a total of 1,000 random data sets, found that tumor location, percentage of solid and mucinous components, presence of Crohn-like response and TIL were strongly associated with the MSI status being present in 976 to 1,000 of the generated models. The growth pattern included in 715 models and the expression of Ki-67 (in 592) and p53 (in 651) were also considered to be predictive of the MSI status and thus included in our optimized model, named RERtest8:
The coefficients of the linear component (x) corresponded to: (a) location (0 = proximal, 1 = distal), (b) growth pattern (0 = expansive, 1 = infiltrative), (c) Crohn-like response (0 = present, 1 = absent), (d) TIL%, (e) solid pattern%, (f) mucinous pattern%, (g) Ki-67%, and (h) p53 accumulation%.
Comparing these data with our original model, it should be noted that TIL has been included in the equation at the expense of the proportion of cribriform pattern which has been left out of the optimized model. Table 4 shows the accuracy, sensitivity, specificity, positive, and negative predicted values achieved by the optimized model. Considering that the expression of Ki-67 and p53 by immunohistochemistry were the variables with a weaker relation to the MSI status and were also the only parameters that could not be determined just by morphology assessment, a second alternative model was constructed that excluded these two variables, RERtest6:
Table 4 Statistical parameters achieved with our optimized models, with and without including immunohistochemistry parameters
| RERTest8 | RERTest6 | |
| Accuracy | 92.48 | 92.09 |
| Sensitivity | 78.14 | 78.01 |
| Specificity | 93.80 | 93.39 |
| Positive predictive value | 53.45 | 51.78 |
| Negative predictive values | 97.93 | 97.91 |
| Total (n) | 615 | 615 |
The coefficients of the linear component (x) corresponded to: (a) location (0 = proximal, 1 = distal), (b) growth pattern (0 = expansive, 1 = infiltrative), (c) Crohn-like response (0 = present, 1 = absent), (d) TIL%, (e) solid pattern%, (f) mucinous pattern%.
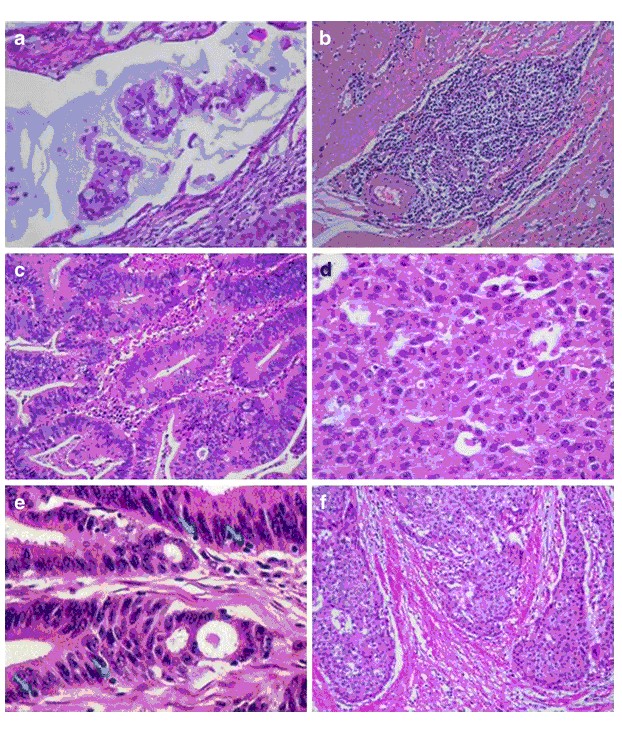
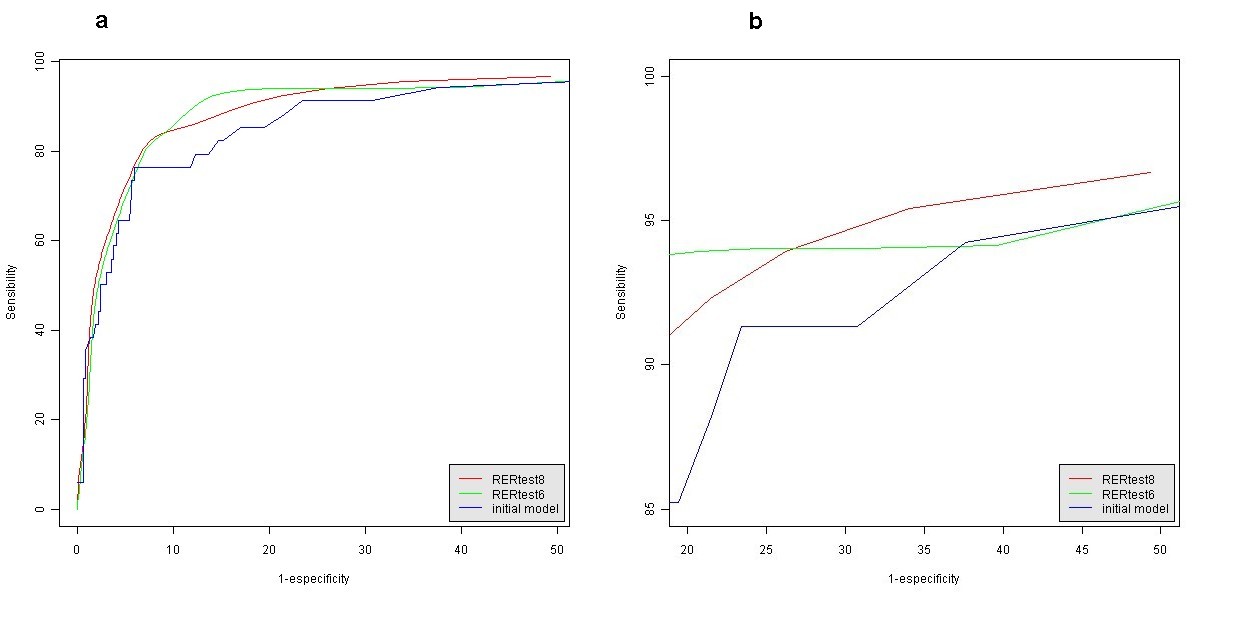
The statistical parameters of this alternative model are also shown in Table 4. A probability value of a tumor being MSS lower than 80% (P < 0.8) was empirically set as a cutoff point to discriminate between MSI-H and MSS tumors with both models. Figure 1 shows representative pictures of the histological parameters finally included in our optimized model RERtest6, and Fig. 2 shows the ROC curves of the optimized models RERtest8 and RERtest6 compared to that of our original model.
Fig. 1 Histological parameters included in our optimized model RERtest6. a Adenocarcinoma with extracellular mucinous pattern (H&E, 400x). b “Crohn like” inflammatory response (H&E, 200x). c Adenocarcinoma without solid pattern (H&E, 200x). d 100% solid pattern (H&E, 400x). e Tumor infiltrating lymphocytes (TIL) (H&E, 400x). f Expansive growth in tumoral margin (H&E, 200x)
Fig. 2 a ROC curves of the optimized and original models and b enlarged view of the area of interest
Discussion
The complexity and cost of the MSI study added to the increasing availability of tailored therapies, and screening programs has triggered the development of models based on pathological features to predict instability status. The model that we had published [15] and has now been further validated, focus on a non-selected population of patients with colorectal cancer aiming at identifying not only possible Lynch syndrome candidates, but any carcinomas exhibiting an MSI-H phenotype and thus a better prognosis.
The multicenter validation carried out with samples from five different centers has proved that our original model was robust enough to overcome not only interobserver variability but also differences between the immunohistochemistry methods employed.
The optimization studies with a global series of 615 colorectal carcinomas gave rise to an improved model, RERtest8, in which the percentage of cribriform structures was left out of the equation while the presence of TIL was included. This new model is in closer concordance with other recently published models where proximal tumor location, presence of TIL, Crohn-like reaction, mucinous component, and poor differentiation were assigned as predictors of MSI status [13, 14]. Nevertheless, there are important differences to be pointed out. The MsPath score published by Jenkins and colleagues [13] looks at the pathology features highlighted by Bethesda guidelines in a series of colorectal carcinomas diagnosed before age 60 with the main aim of identifying Lynch syndrome candidates. The study published by Greenson and colleagues [14], even though it does not restrict its series by the patient’s age, finds age under 50 to be the only independent predictor of MSI-H status and includes this parameter as the second most influencing after TIL in the equation of their Pathscore. In contrast, we have not found age to be predictive of the MSI status neither in our initial multivariate logistic analysis nor in our current optimized model. This may be due to the high mean age of patients in our series 69.2. This high mean age could also be an explanation for the low percentage of MSI-H cases found in our study (8.5%), compared to other series described in literature (10–15%), [1] since HNPCC patients typically have an early onset of the illness.
The impact of each of the parameters included also varies between models. Proximal tumor location and presence of peritumoral Crohn-like inflammatory response are the strongest predictive parameters in our RERtest8 model followed by expansive growth pattern and TIL. The expansive growth pattern was not included in the other above-mentioned models, although it was found to be a MSI predictor by univariate analysis in the study of Greenson. The grade of differentiation was assessed and integrated in our model by looking at the percentage of solid component (Fig. 1c, d). The presence of mucinous pattern was also measured as a percentage in order to better adjust its influence, since the mean of the mucinous pattern found in our MSI-H group was as low as 32.3%. The two immunohistochemistry parameters, p53 expression and Ki-67 proliferative index, were the weakest factors incorporated in our RERtest8, this together with the fact that they are the only parameters that are not morphological allowed us to reconstruct our model excluding them, the RERtest6 model.
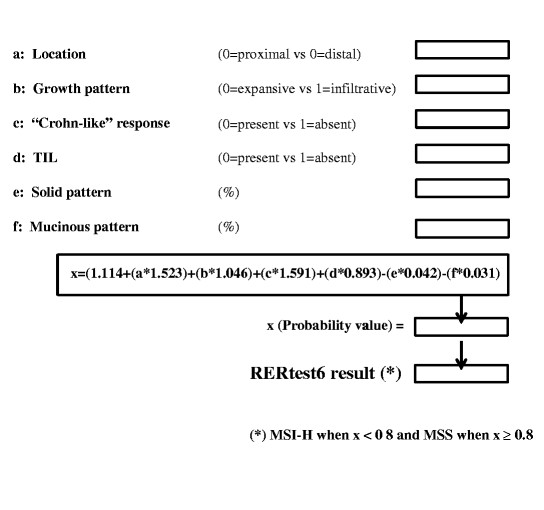
The general approach of our optimization studies was to maximize the negative predictive value of our models in order to construct a tool that would allow the reduction of cases to be tested for MSI and could be easily implemented in an Excel document for daily use (Fig. 3). In this way, we present two models RERtest8 and RERtest6, with and without immunohistochemistry parameters, with negative predictive values of 97.9% that allow the reduction of the cases to be tested for MSI to approximately 10%; we proposed that only those cases predicted as MSI-H should be confirmed as such. Furthermore, the easy evaluation of the parameters included in the models, the fact that it is not influenced by interobserver variability and the possibility of excluding immunohistochemistry parameters, renders it a very useful tool for the routine practice and can reinforce the current clinical protocols to detect the subset of colorectal cancer patients bearing MSI-H phenotype, and thus a better prognosis, and HNPCC risk.
Fig. 3 RERtest6 Model in Microsoft’s Excel format. As shown, the pathologist assessing the case should include either 0 or 1 in the cells corresponding to categorical variables (a–d) and the percentage of solid or mucinous component in cells e and f. The model’s formula result is calculated in a separate cell, and in the last cell the case is classified as MSS or MSI-H depending on the x value
________________________________________
Acknowledgment The authors thank Eva Torija from BIOPAT for her secretarial assistance in data collection.
Conflict of interest statement We declare that we have no conflict of interest.
________________________________________
References
- Ogino S, Goel A (2008) Molecular classification and correlates in colorectal cancer. J Mol Diagnostics 10:13–27, s.l
- Gryfe RH, Kim H, Hsieh ET (2000) Tumor microsatellite instability and clinical outcome in young patients with colorectal cancer. N Engl J Med 342:69–77, s.l
- Fearon ER, Volgestein B (1990) A generic model for colorectal tumorigenesis. Cell 61:759–767
- Thibodeau SN, Bren G, Schaid D (1993) Microsatellite instability in cancer of proximal colon. Science 260:816–819, s.l
- Jass JR, Do KA, Simms LA (1998) Morphology of sporadic colorectal cancer with DNA replication errors. Gut 42:673–679, s.l
- Fujiwara T, Stolker JM, Watanabe T et al (1998) Accumulated clonal genetic alterations in familial and sporadic colorectal carcinomas with widespread instability in microsatellite sequences. Am J Pathol 153:1063–1078, s.l
- Herman JG, Umar A, Polyak K et al (1998) Incidence and functional consequences of hMLH1 promoter hypermethylation in colorectal carcinoma. Proc Natl Acad Sci 95:6870–6875, s.l
- Arnold CN, Goel A, Boland CR (2003) Role of hMLH1 promoter hypermethylation in drug resistance to 5-fluorouracil in colorectal cancer cell lines. Int J Cancer 106:66–73, s.l
- Ribic CM, Sargent DJ, Moore MJ et al (2003) Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med 349:247–257
- Jass JR (2004) HNPCC and sporadic MSI-H colorectal cancer: a review of the morphological similarities and differences. Familial Cancer 3:93–100, s.l
- Kakar S, Aksoy S, Burgart LJ, Smyrk TC (2004) Mucinous carcinoma of the colon: correlation of loss of mismatch repair enzymes with clinicopathologic features and survival. Mod Pathol 17:696–700
- Ogino S, Brahmandam M, Cantor M et al (2006) Distinct molecular features of colorectal carcinoma with signet ring cell component and colorectal carcinoma with mucinous component. Mod Pathol 19:59–68, s.l
- Jenkins MA, Hayashi S, O’Shea et al (2007) Pathology features in Bethesda guidelines predict colorectal cancer microsatellite instability: a population-based study. Gastroenterology 133:48–56
- Greenson JK, Huang S, Herron C et al (2009) Pathologic predictors of mirosatellite instability in colorectal cancer. Am J Surg Pathol 33:126–133
- Colomer A, Erill N, Vidal A et al (2005) A novel logistic model based on clinicopathological features predicts microsatellite instability in colorectal carcinomas. Diagn Mol Pathol 14:213–223
- Michael-Robinson JM, Biemer-Huttmann A, Purdie DM et al (2001) Tumour infiltrating lymphocytes and apoptosis are independent features in colorectal cancer stratified according to microsatellite instability status. Gut 48:360–366
- Hamilton SR, Aaltonen LA (2000) Pathology and genetics of tumours of the digestive system. eds World Health Organization Classification of Tumors, IARC Press, Lyon, pp 103–143
- Boland CR, Thibodeau SN, Hamilton SR et al (1998) A National Cancer Institute Workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 58:5248–5257
- Jen J, Kim H, Piantadosi S et al (1994) Allelic loss of chromosome 18q and prognosis in colorectal cancer. N Engl J Med 331:213–221
- Jones MH, Nakamura Y (1992) Detection of loss of heterozygosity at the human TP53 locus using a dinucleotide repeat polymorphism. Genes Chromosom Cancer 5:89–90
- Friedman J, Hastie T, Tibshirani R (2008) Glmnet: Lasso and elastic-net regularized generalized linear models: http://www-stat.stanford.edu/∼hastie/Papers/glmnet.pdf
- Friedman J, Hastie T, Höfling H, Tibshirani R (2007) Pathwise coordinate optimization. Ann Appl Stat 1(2):302–332
- R Development Core Team. R (2008) A language and environment for statistical computing. s.l.: R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org
- Schwarz G (1978) Estimating the dimension of a model. Ann Stat 6:461–464